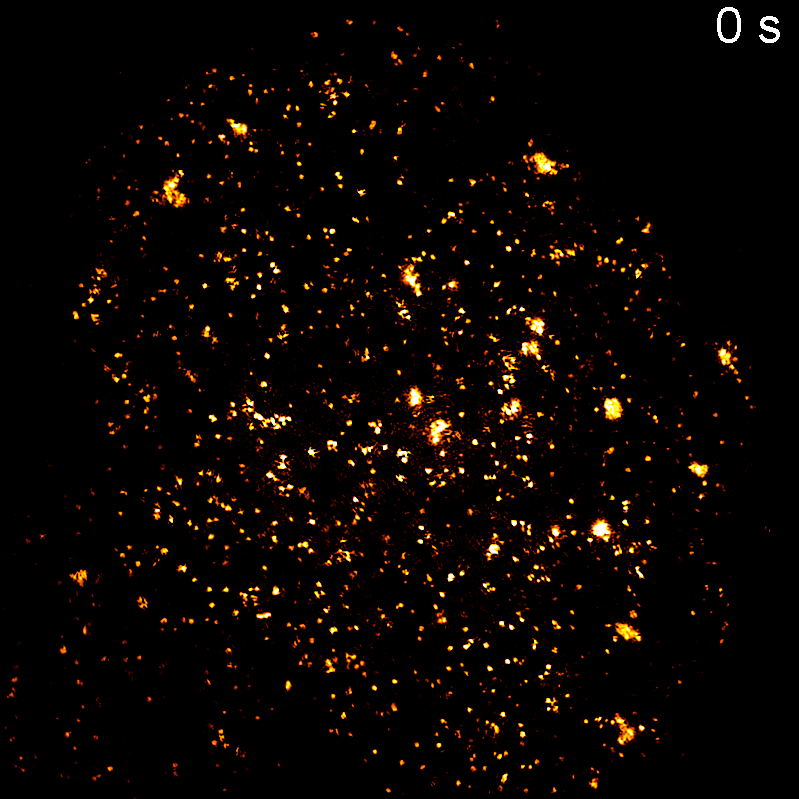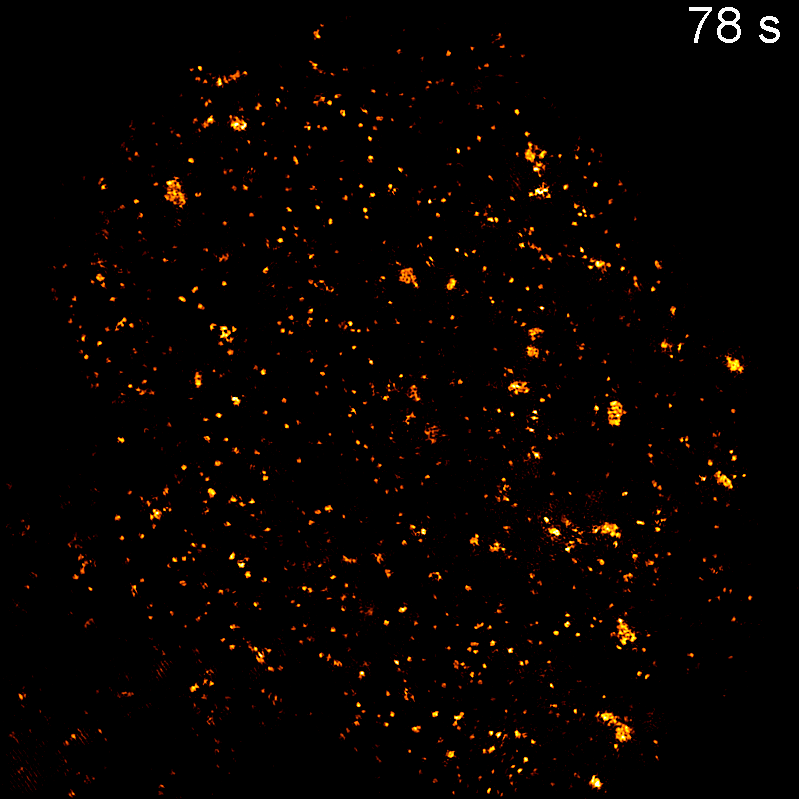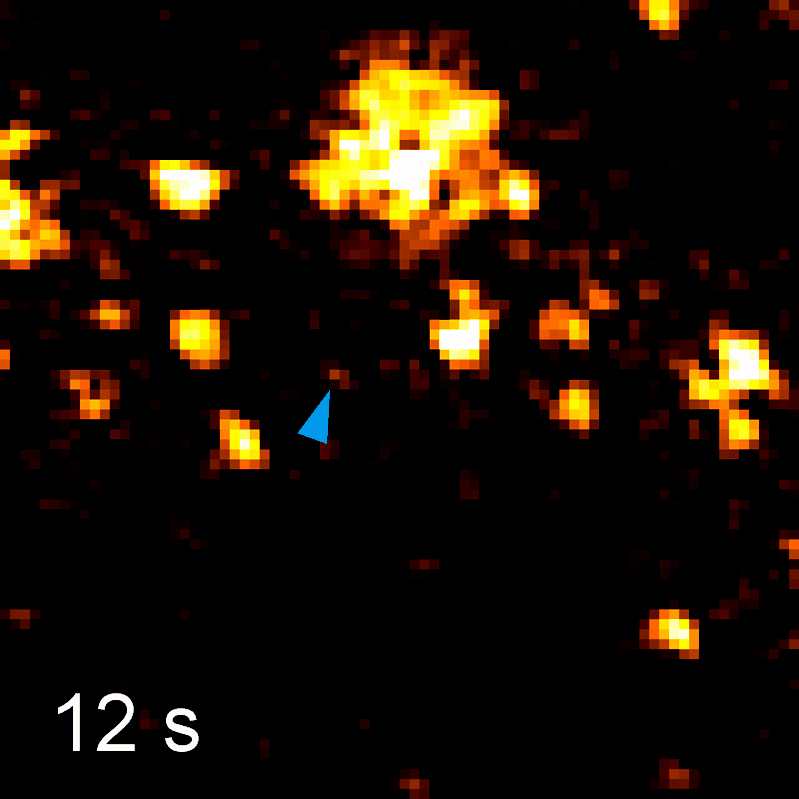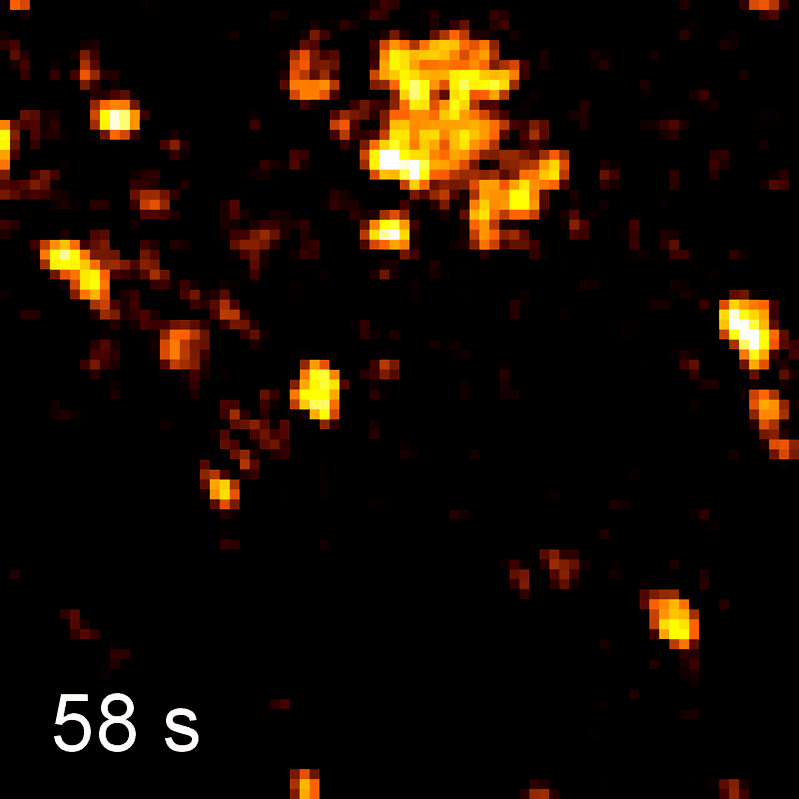
Endocytosis
Membrane trafficking
Quantitative studies of clathrin-mediated endocytosis and intracellular dynamics.
Intracellular Dynamics Laboratory
We build microscopes and computational tools to watch cells in real time — studying how membranes bend, how cargo gets internalized, and how mechanics shape disease.
01 - Highlights
Visual highlights spanning endocytosis, mechanobiology, and imaging innovation. These are the systems and questions that drive our day-to-day science.
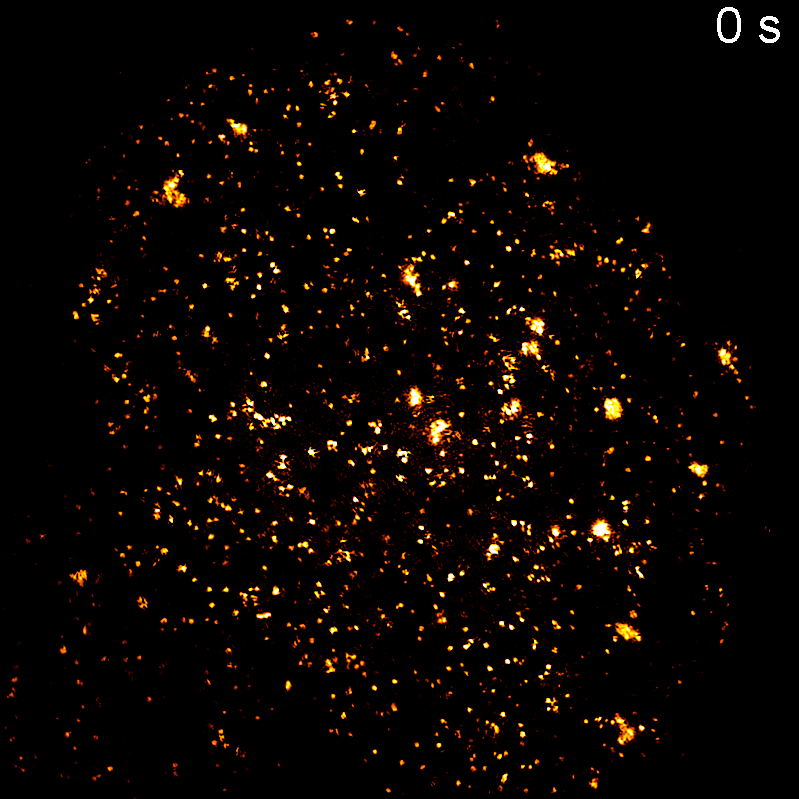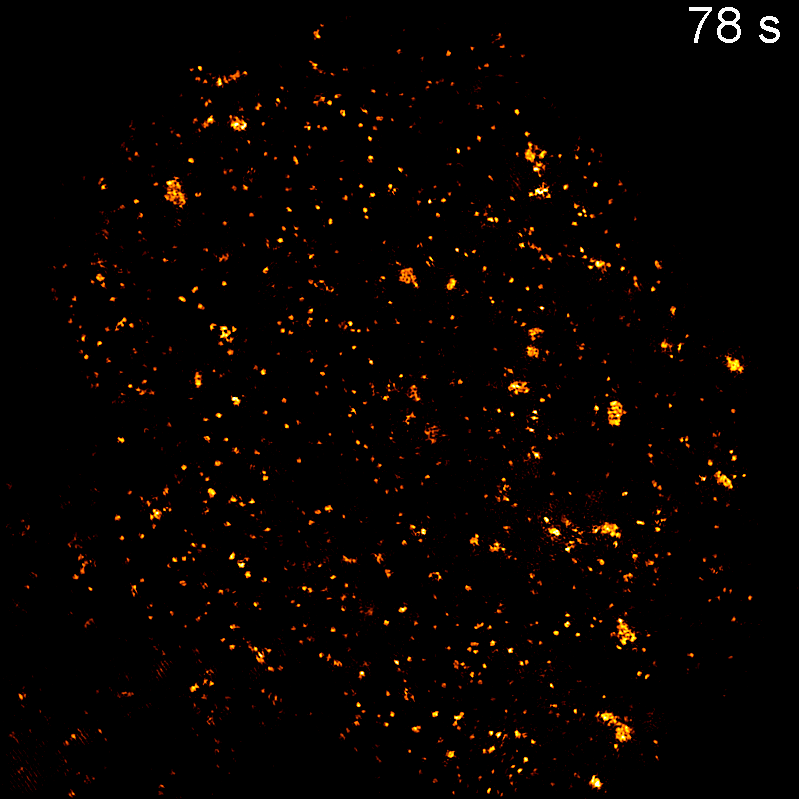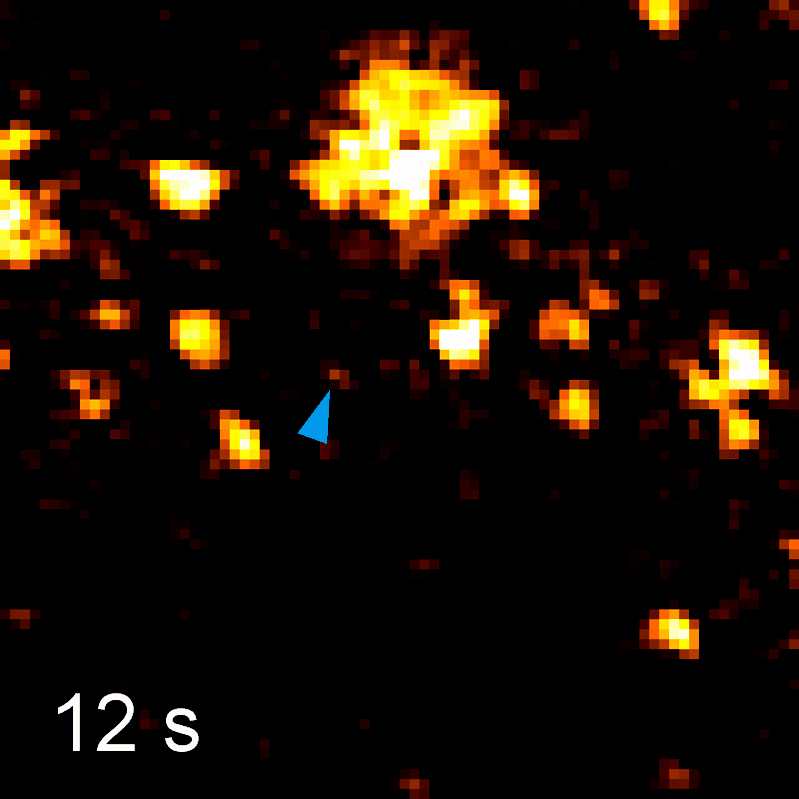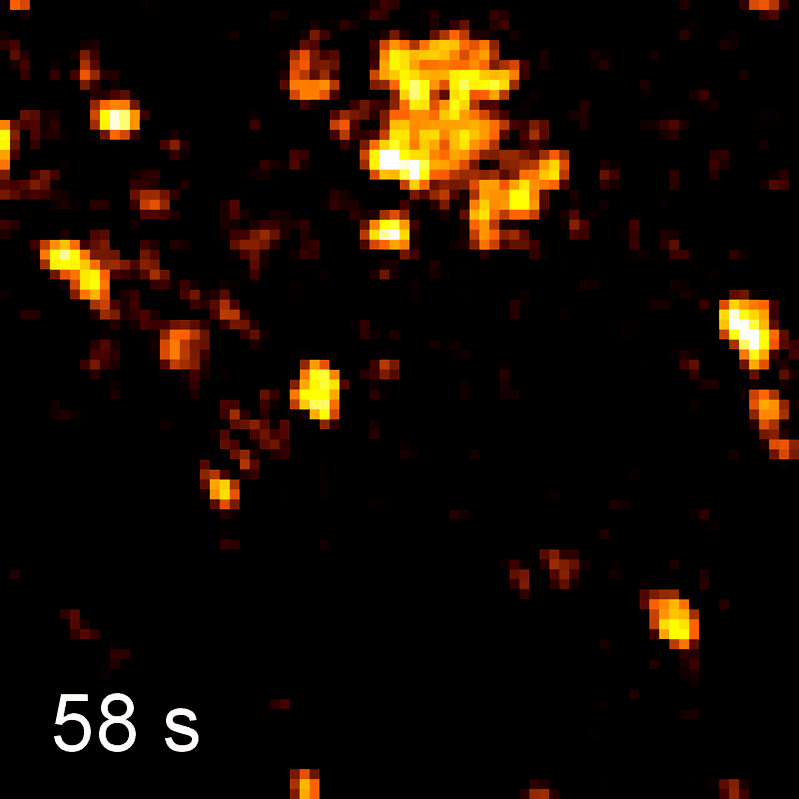
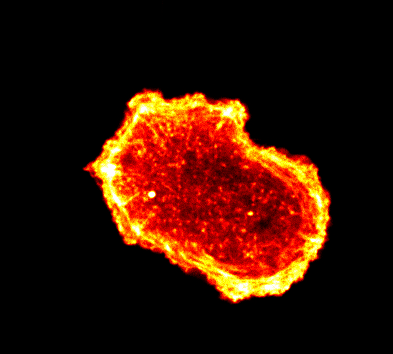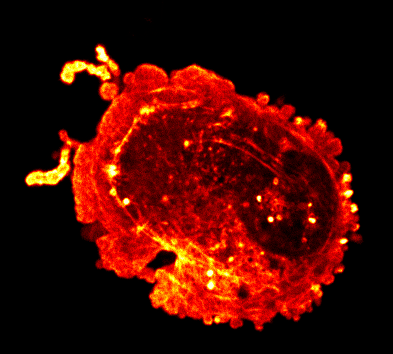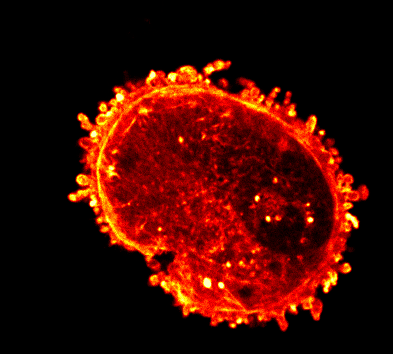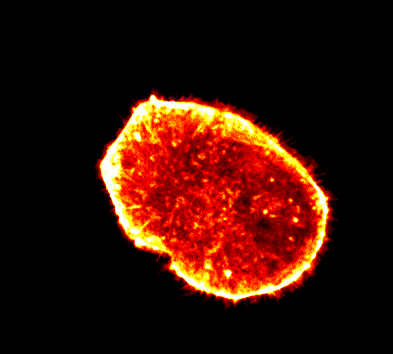
Endocytosis
Quantitative studies of clathrin-mediated endocytosis and intracellular dynamics.
Disease
How cell mechanics influence signaling, apoptosis, and cancer-related phenotypes.

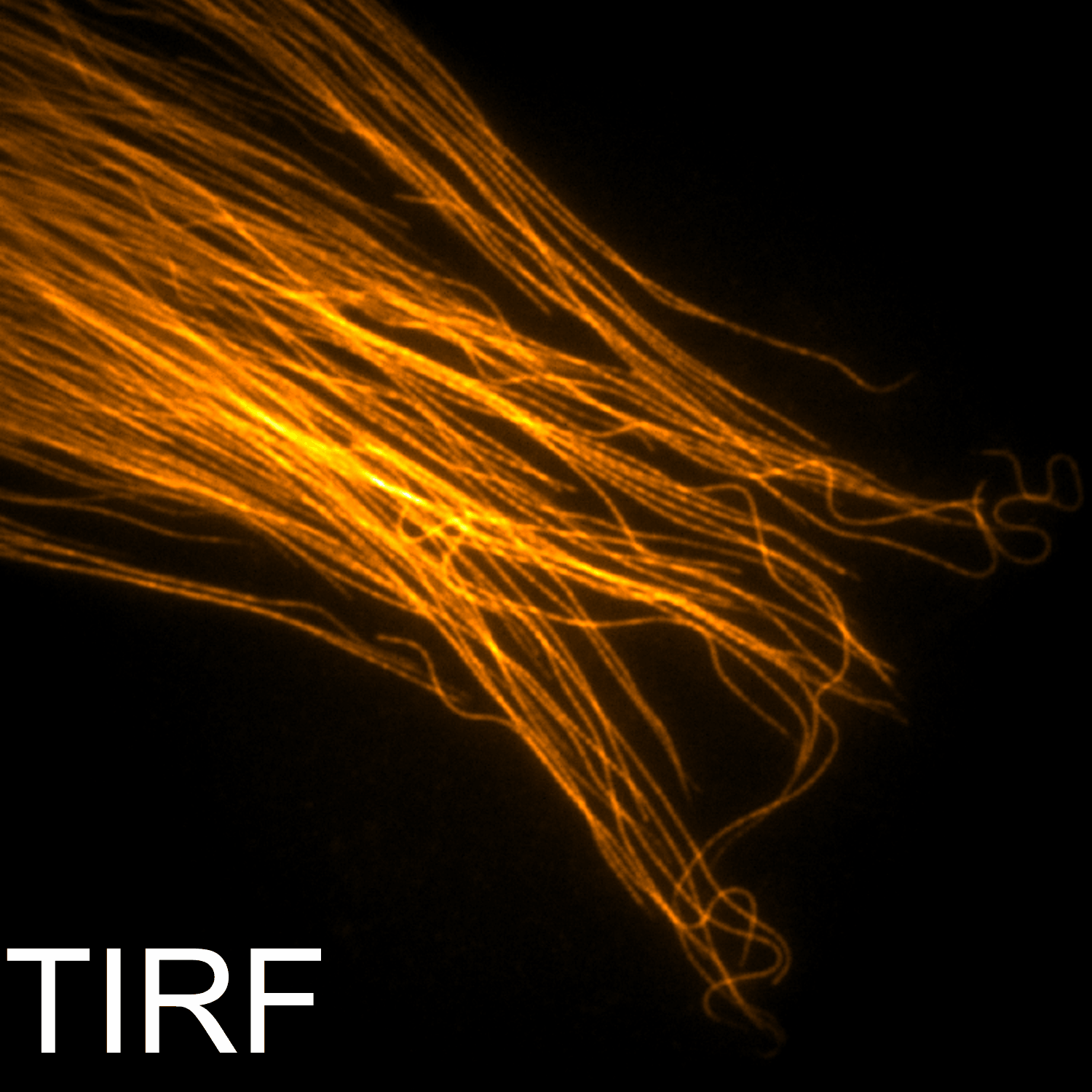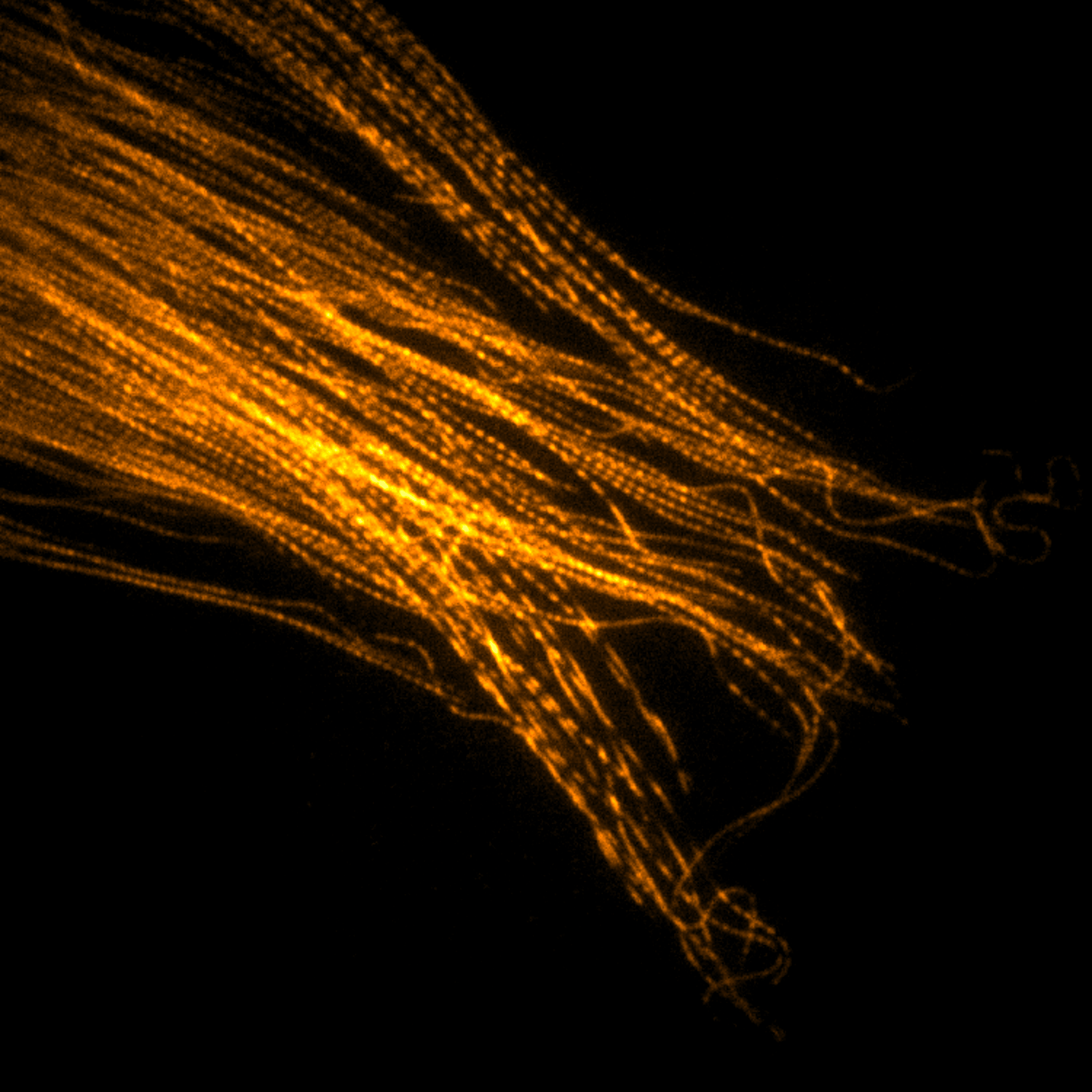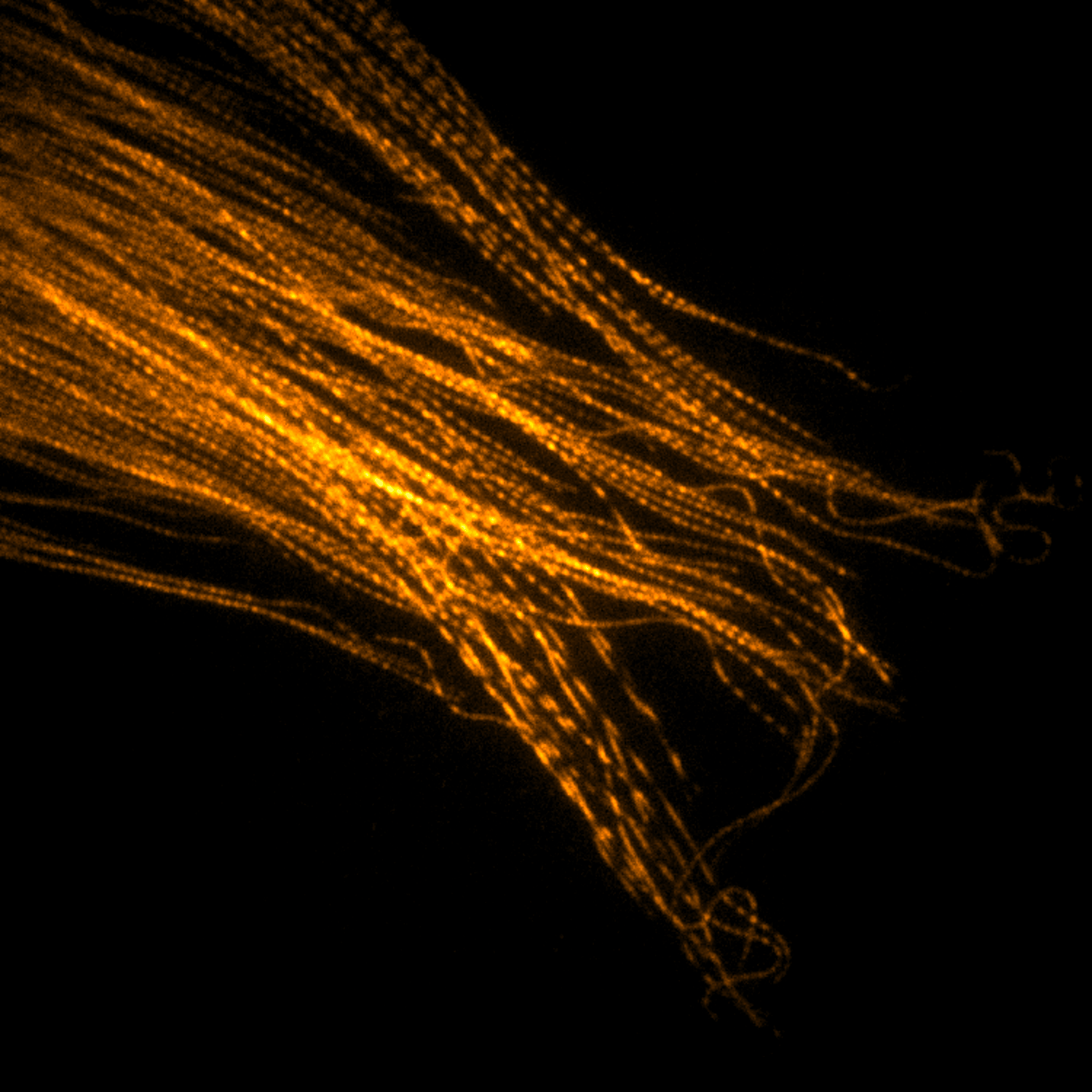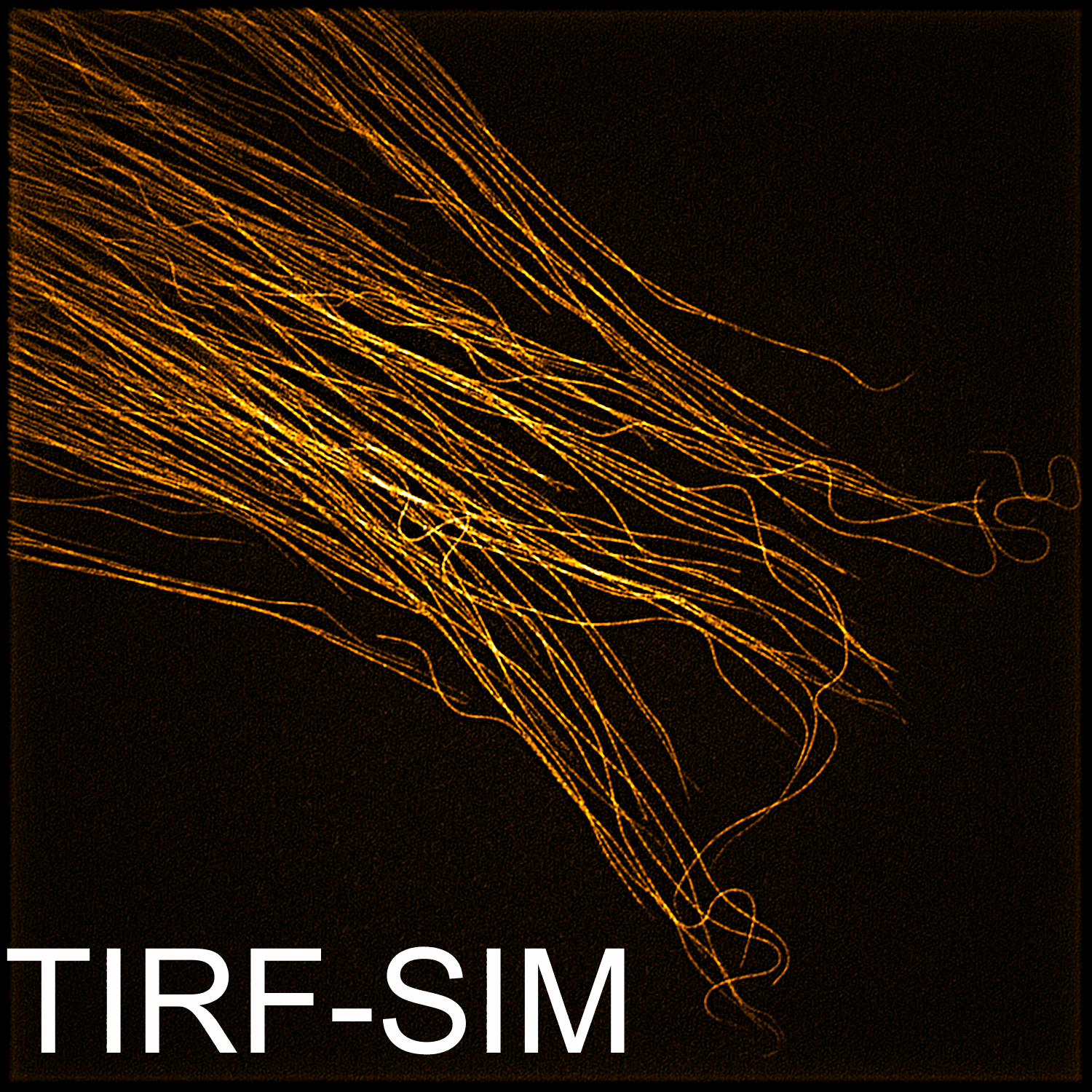
Imaging
Instrumentation and computational methods for super-resolution and live-cell imaging.

Structure
How coated pits and plaques form, curve, and internalize.
Computation
Image analysis pipelines that reveal dynamic intracellular organization.
People
An interdisciplinary group at the interface of physics and cell biology.
02 - Research
Our work links instrumentation, computation, and translation. We aim to uncover how molecular-scale events give rise to emergent cellular behaviors and how that knowledge can be turned toward therapy.
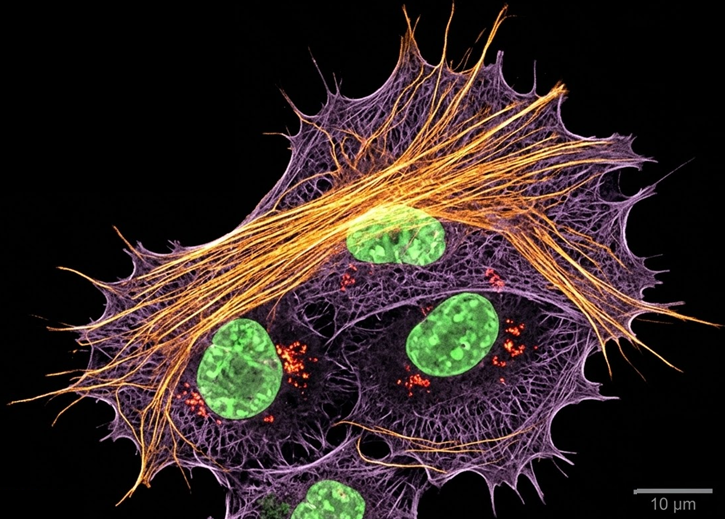
2.1
We develop and apply fluorescence microscopy approaches that resolve intracellular dynamics at high spatial and temporal resolution. Techniques include TIRF microscopy, structured illumination microscopy, and variable-angle illumination — selectively visualizing membrane-proximal processes like clathrin-mediated endocytosis.
We pair instrumentation with computational imaging that incorporates deep learning, temporal information, and three-dimensional data to extract super-resolution insight from live-cell datasets, bridging the gap between high-resolution imaging and fast cellular dynamics.
By integrating optical instrumentation, quantitative analysis, and machine learning, we aim to uncover how molecular-scale events give rise to emergent cellular behaviors in space and time.
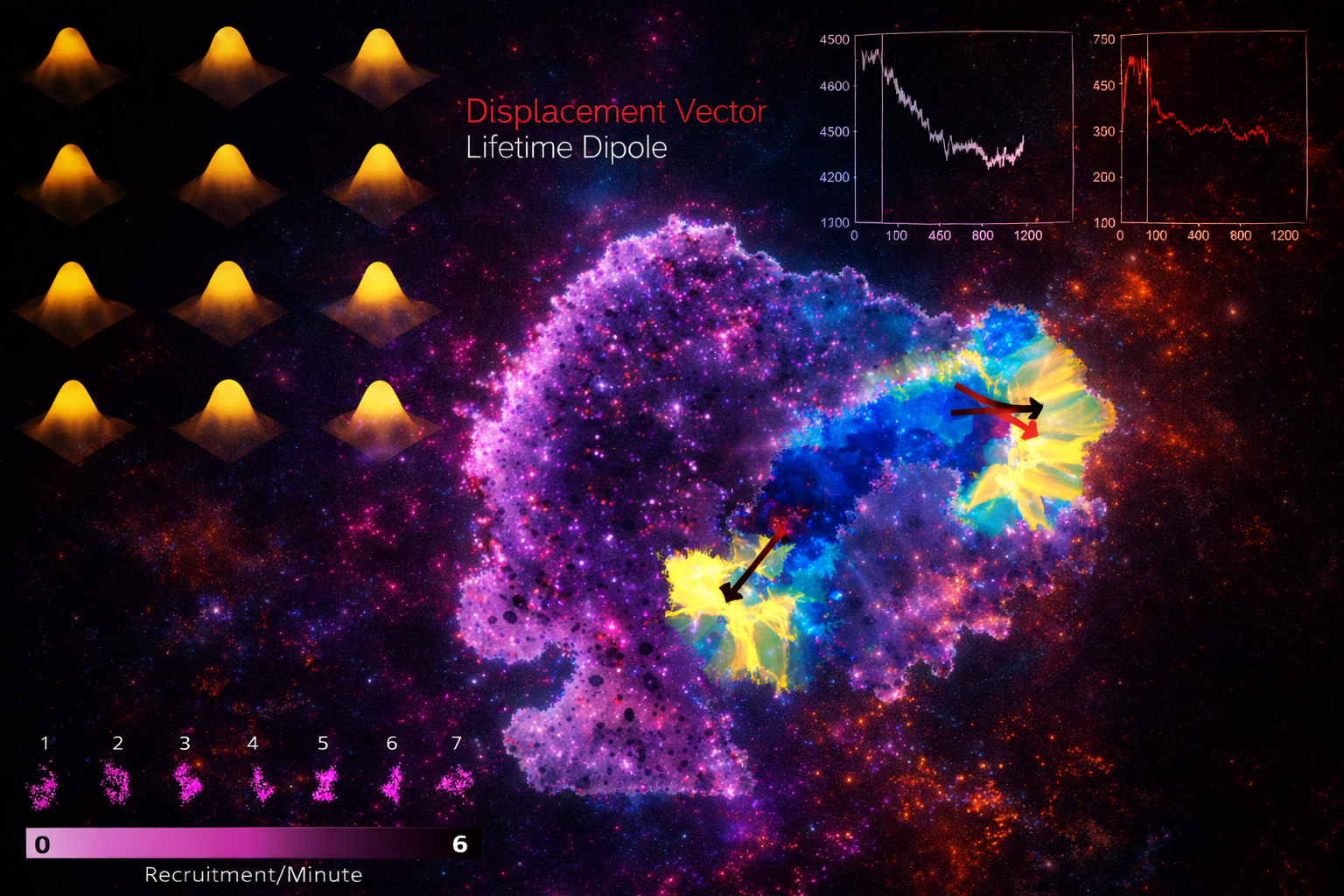
2.2
Because intracellular processes are dynamic and heterogeneous, we build analysis frameworks that move beyond descriptive imaging toward rigorous quantitative inference. Our methods include particle detection and tracking, intensity-based growth-rate analysis, and trajectory classification.
By combining experimental measurements with analytical and computational models, we link observed fluorescence signals to molecular processes - adaptor recruitment, curvature generation, vesicle formation.
Overall, our goal is to transform rich imaging datasets into predictive, mechanistic understanding of cellular processes.
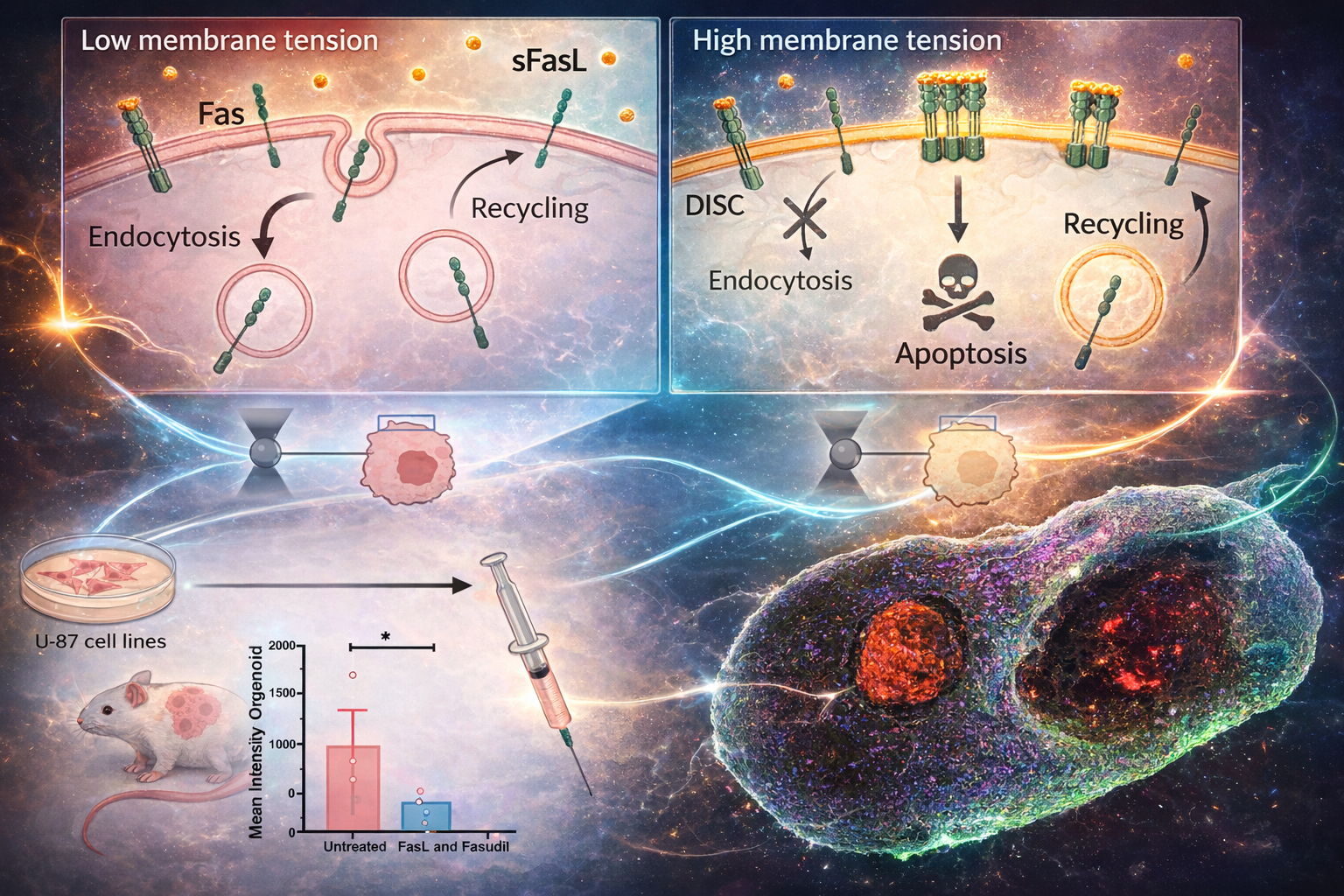
2.3
We explore how cellular mechanics and membrane trafficking can be leveraged to improve therapeutic outcomes — particularly in cancer. The physical state of a cell, its stiffness, membrane tension, and trafficking activity, directly influences how it responds to external signals.
We have shown that perturbing endocytic pathways can sensitize cancer cells to apoptosis. Building on this, we investigate small-molecule "mechanosensitizers" that shift the physical and trafficking properties of tumor cells to make them more vulnerable to immune-mediated killing.
Our approach integrates biophysics, live-cell imaging, and translational research to identify strategies that complement existing therapies such as immunotherapy. The long-term goal is to develop new treatment paradigms where tuning the mechanical and trafficking state of cells enhances therapeutic efficacy while maintaining safety.
03 - Members
An interdisciplinary group spanning physics, biology, and computation.

Illustration of the Kural Lab team, created by Emily Chan.

Comert Kural, Ph.D.
Associate Professor, Department of Physics · Biophysics Graduate Program · Molecular, Cellular & Developmental Biology Program

Emily Chan
B.A. Macalester College - Chemistry, Physics minor (2019)

Tianyao Wu
B.Sc. University of Wisconsin-Madison - Physics (2015)

Cris Thompson
B.Sc. Bates College - Physics (2019)

Valeria Arteaga Muniz
B.Sc. University of Texas at El Paso - Physics (2022)

Aritra Mondal
M.Sc. IISER Kolkata - Biological & Physical Sciences (2022)

Hermes Hermes

Arvin Alam

Henry Jiang

Hirak Basu
Duke Medical Physics

Umida Djakbarova, Ph.D.
Arcus Biosciences

Marlin Keller
UW-Madison Medical Physics

Caleb Smith
Duke Medical Physics

Connor Luellen
UC Berkeley Biophysics

Yasaman Madraki
Roche

Ata Akatay
National Metrology Institute of Türkiye

Jama Hersi

Lauren Riede

Hoda Akl, M.Sc.

Nathan Willy, Ph.D.

Salih Silahli, Ph.D.

Scott Huber, Ph.D.

Joshua P. Ferguson, Ph.D.

Ali Adali, Ph.D.

Farah Hasan

Spencer P. Heidotting

Matthew Webber

Daniel Hoying
Esra Aygun
Sevde Goker
Tugba Atabey
Vannimul Hem
04 - Software
MATLAB tools we have developed and released for the community. All sources are downloadable below.
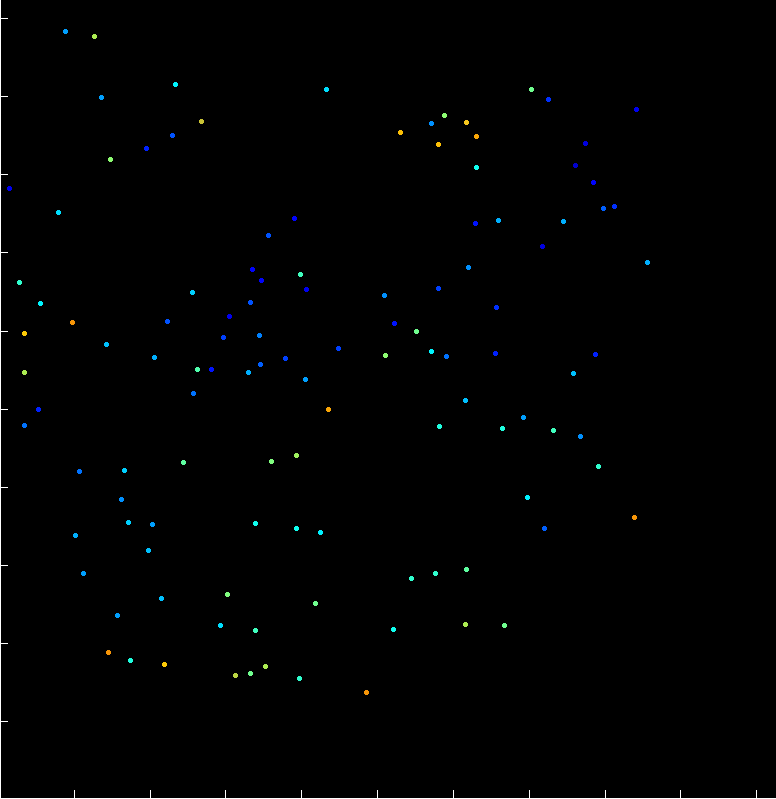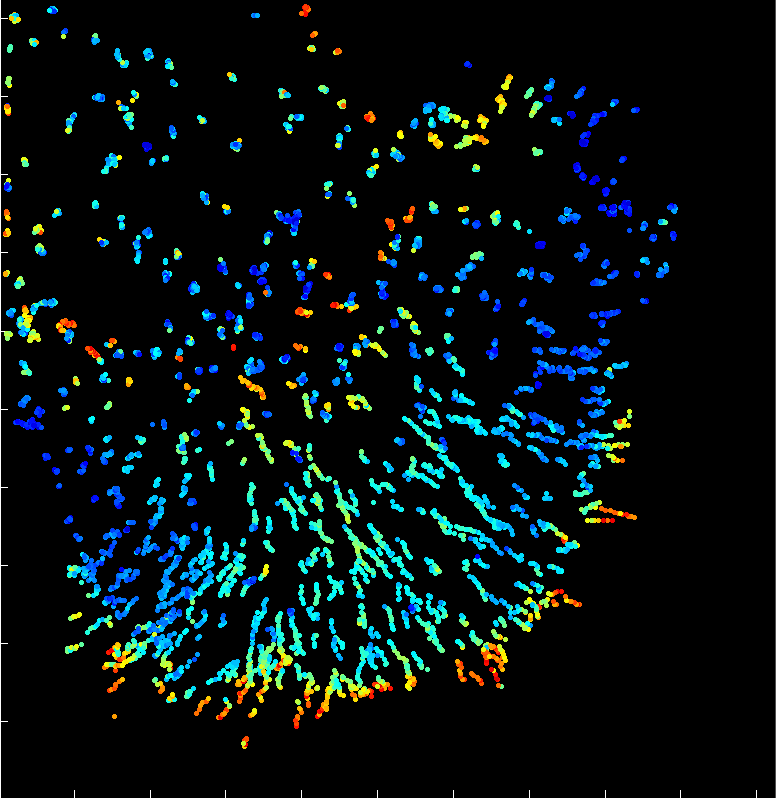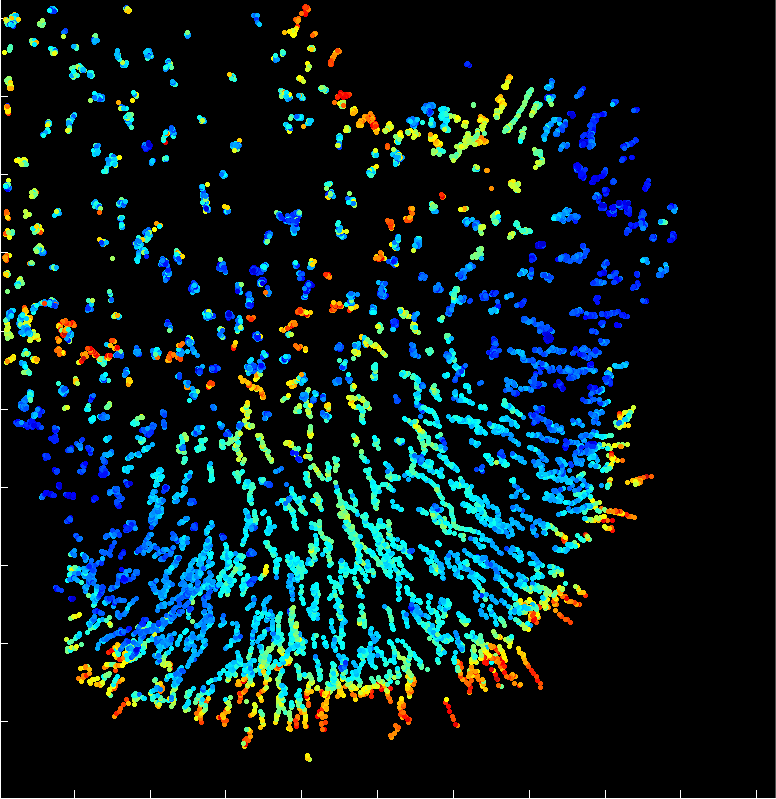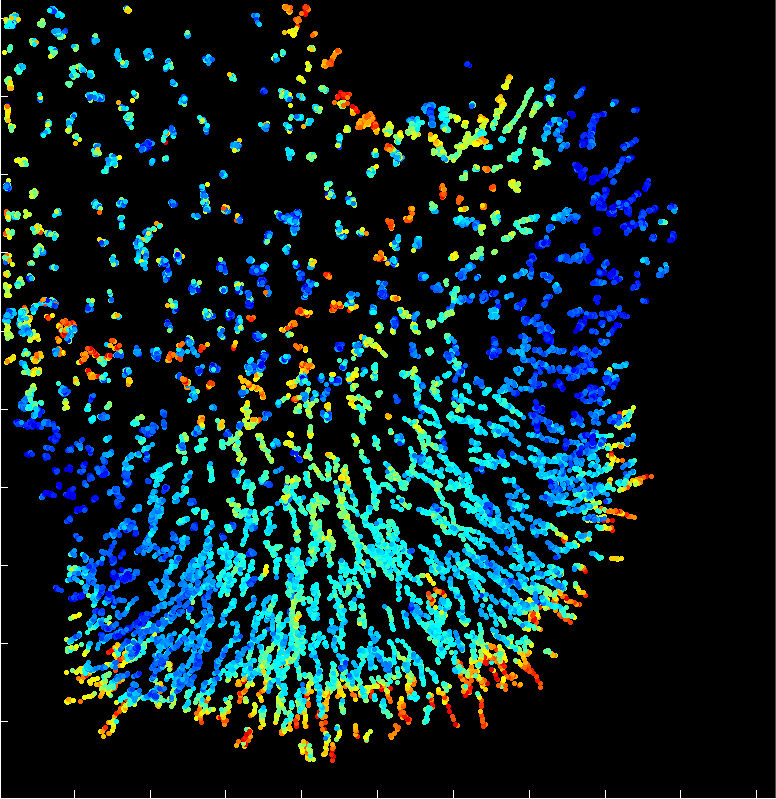
A fast two-dimensional particle tracking program. Detects fluorescent spots using a threshold determined over a Mexican-hat filtered image, then connects maxima in time by linking mutually nearest neighbors. Input: a 2D multipage TIFF. Output: a MAT file of tracked positions and intensities.
Download TraCKer.m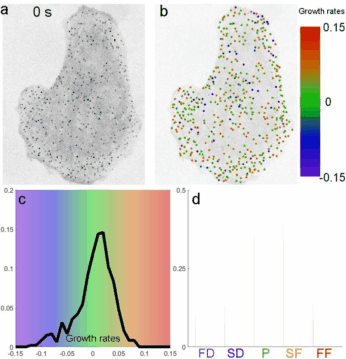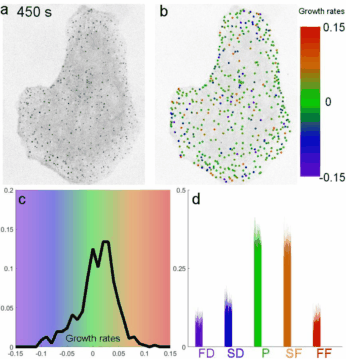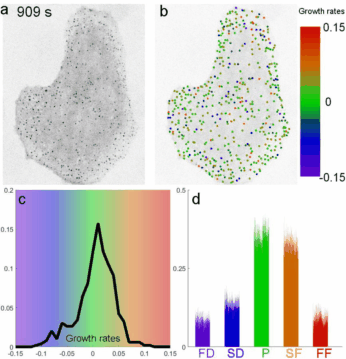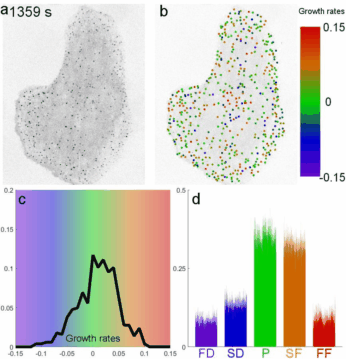
Determines clathrin coat growth-rate distributions from intensity traces. Takes the TraCKer intensity output, the movie frame rate, and a global background value (signal with SNR = 1). Outputs a cell array of normalized slope values.
Download slope_finding.m
createTraceLibrary groups clathrin coat intensity traces into
clusters that share similar trace lengths and intensity profiles. Similarity
is judged using trace_dist. Each cluster yields an average trace
and growth-rate histogram for use as a comparison library.
05 - Publications
A selection of recent peer-reviewed work from the lab. Older publications are listed in the expandable archive at the end.
Accepted 2026
Accepted 2026
Accepted 2026
Journal of Chemical Physics, 163, 151101 2025
Bioengineering, 12(11):1148 2025
NPJ Imaging, 3(1):5 2025
Biochemical Society Transactions, 52(4):1703-1713 2024
Cell Death & Disease, 15(6):440 2024
Frontiers in Molecular Biosciences, 21(9):959737 2022
Developmental Cell, 56(22):3146-59 2021
PNAS, 118(25):e2010438118 2021
Biology of the Cell, 113(8):344-373 2021
Nature Methods, 16(1):103-110 2019
Journal of Cell Science, 130:3611-3617 2017
Molecular Biology of the Cell, 28(24):3480-3488 2017
Journal of Cell Biology, 214(3):347-58 2016
Small, 12(3):308-20 2016
mBio, 6(6):e01541-15 2015
Molecular Biology of the Cell, 26(11):2044-53 2015
Molecular Biology of the Cell, 24(8):1196-207 2013
Cell Reports, 2(5):1111-1119 2012
Methods in Enzymology, 505:59-80 2012
Nature Cell Biology, 13(9):1124-31 2011
Methods in Enzymology, 475:1-26 2010
Biochemistry, 48(22):4663-5 2009
PNAS, 105(29):10011-6 2008
PNAS, 104(13):5378-82 2007
Journal of Cell Biology, 176(5):641-51 2007
Journal of Physics: Condensed Matter, 17:S3979-95 2005
Science, 308(5727):1469-72 2005
Journal of Optics A: Pure and Applied Optics, 3:184-9 2001
06 - News
Awards, new publications, milestones, and instrument arrivals.
2025

Valeria Arteaga-Muniz received the Molecular Biophysics Training Program best oral presentation award. Congratulations Valeria.

Emily Chan received the Biophysics Graduate Program best oral presentation award. Congratulations Emily.

Approaching Maximum Resolution in Structured Illumination Microscopy Via Accurate Noise Modeling is now accepted for publication by NPJ Imaging.
2024
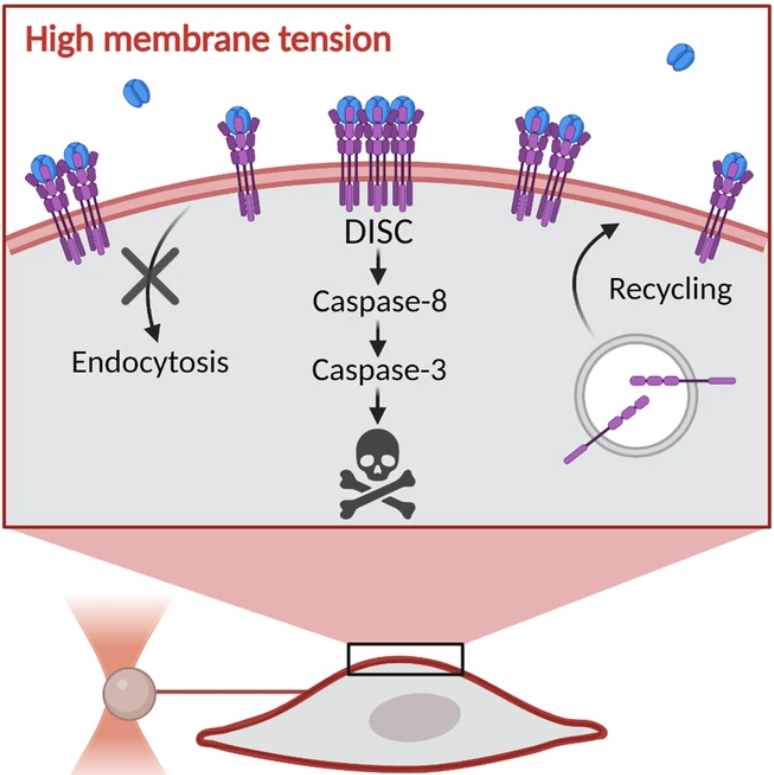
Mechano-inhibition of Endocytosis Sensitizes Cancer Cells to Fas-induced Apoptosis is now accepted for publication in Cell Death and Disease.
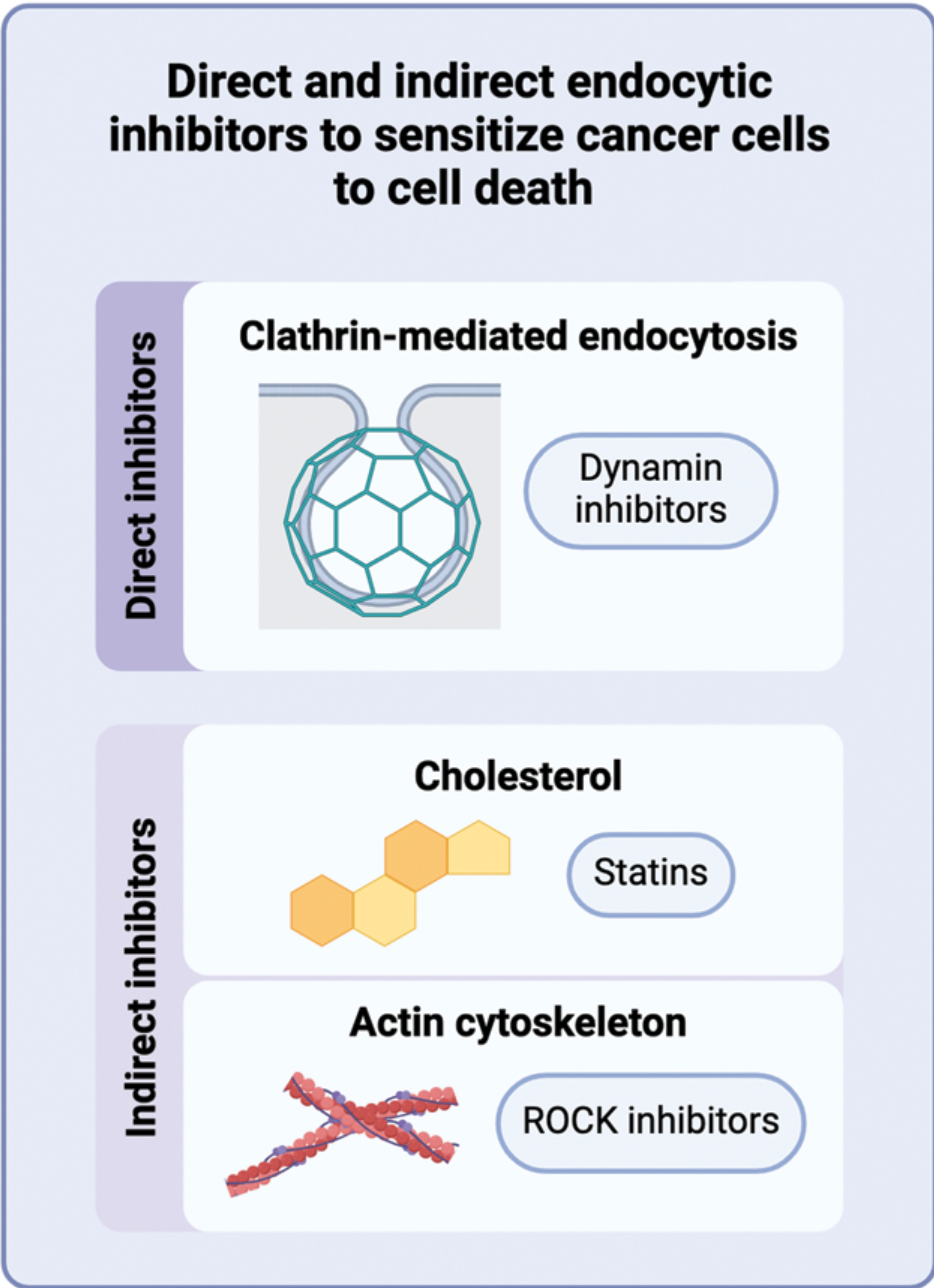
Targeting endocytosis to sensitize cancer cells to programmed cell death is now accepted for publication in Biochemical Society Transactions.
2023

Umida Djakbarova has been awarded a Pelotonia Scholars Symposium Award. Congratulations Umida.

Cris Thompson has been awarded a National Society of Black Physicists (NSBP) best poster award. Congratulations Cris.
2022

Emily Chan has been awarded a Biophysical Society Travel Award. Congratulations Emily.

We have raised $1,900 for Pelotonia. Good job team.
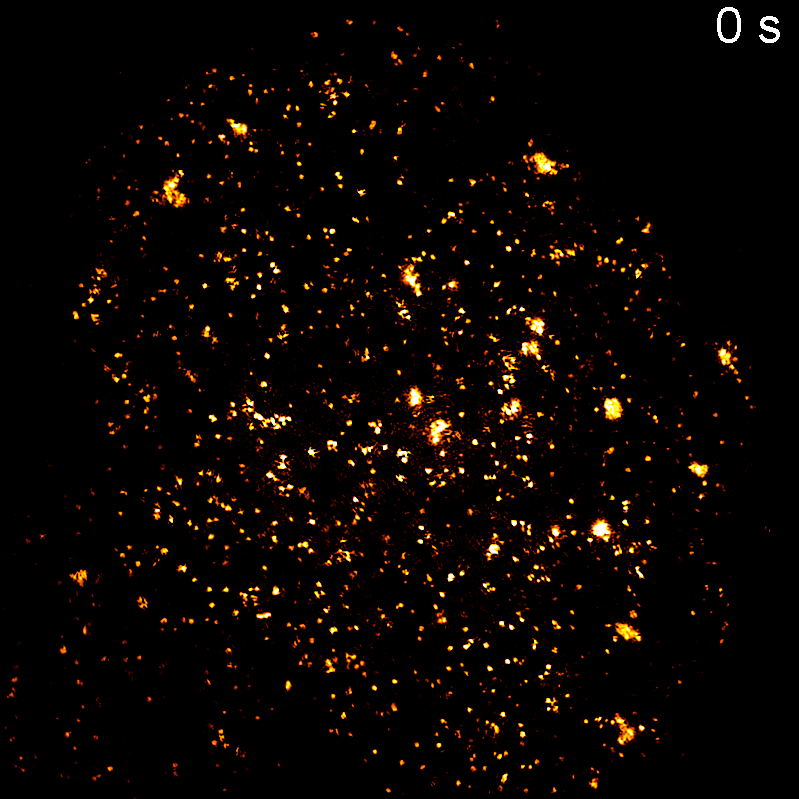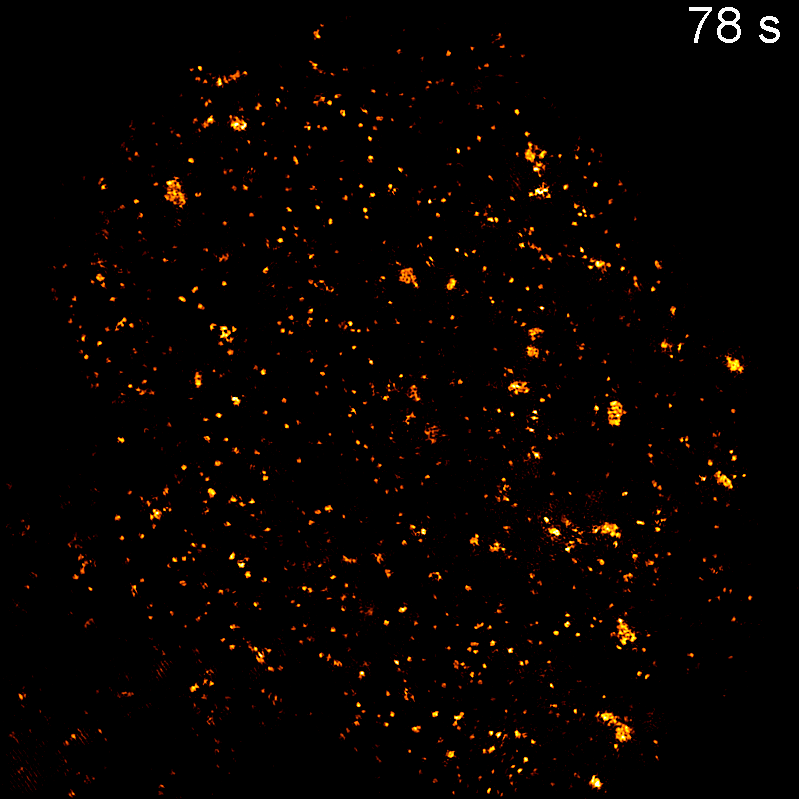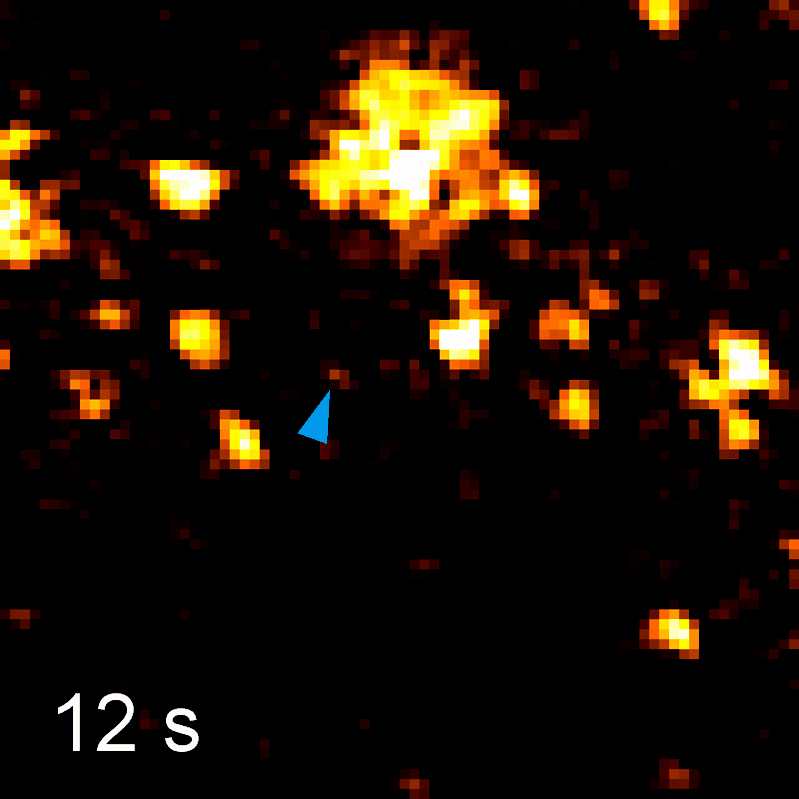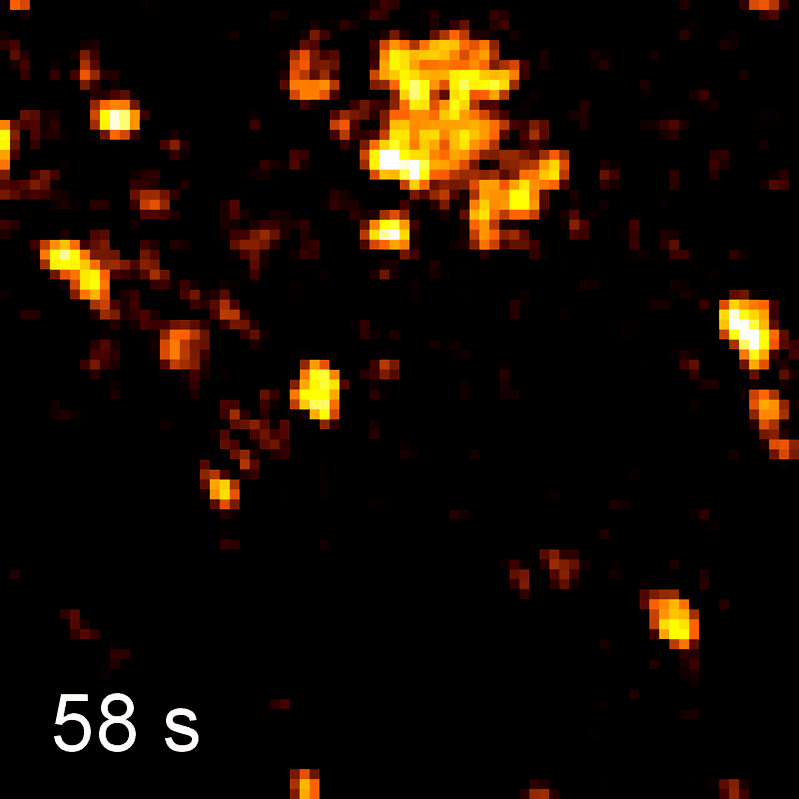
Endocytosis at Extremes: Formation and Internalization of Giant Clathrin-coated Pits Under Elevated Membrane Tension is now accepted for publication by Frontiers Molecular Biosciences.

Our Dual-view Inverted Selective Plane Illumination Microscope (diSPIM) is up and running.

Emily Chan has been awarded a Pelotonia Doctoral Fellowship. Congratulations Emily.
2021


De novo Endocytic Clathrin Coats Develop Curvature at Early Stages of Their Formation is now accepted for publication by Developmental Cell.

Farewell lunch for Connor. We wish him all the best at UC Berkeley.

We have raised $2,800 for Pelotonia. Good job team.
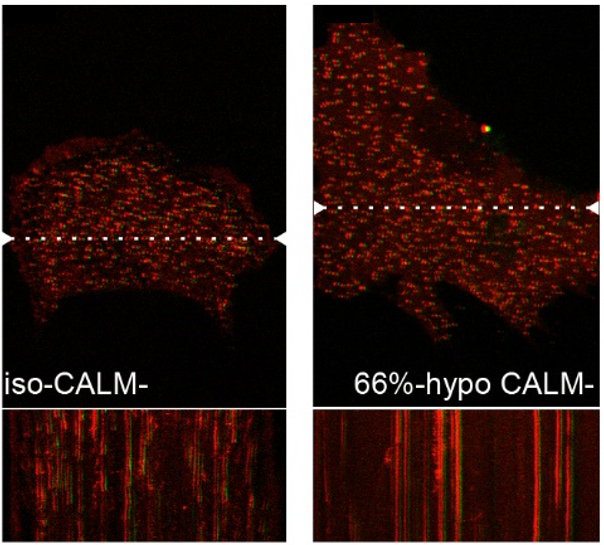
CALM supports clathrin-coated vesicle completion upon membrane tension increase is published in Proceedings of the National Academy of Sciences.
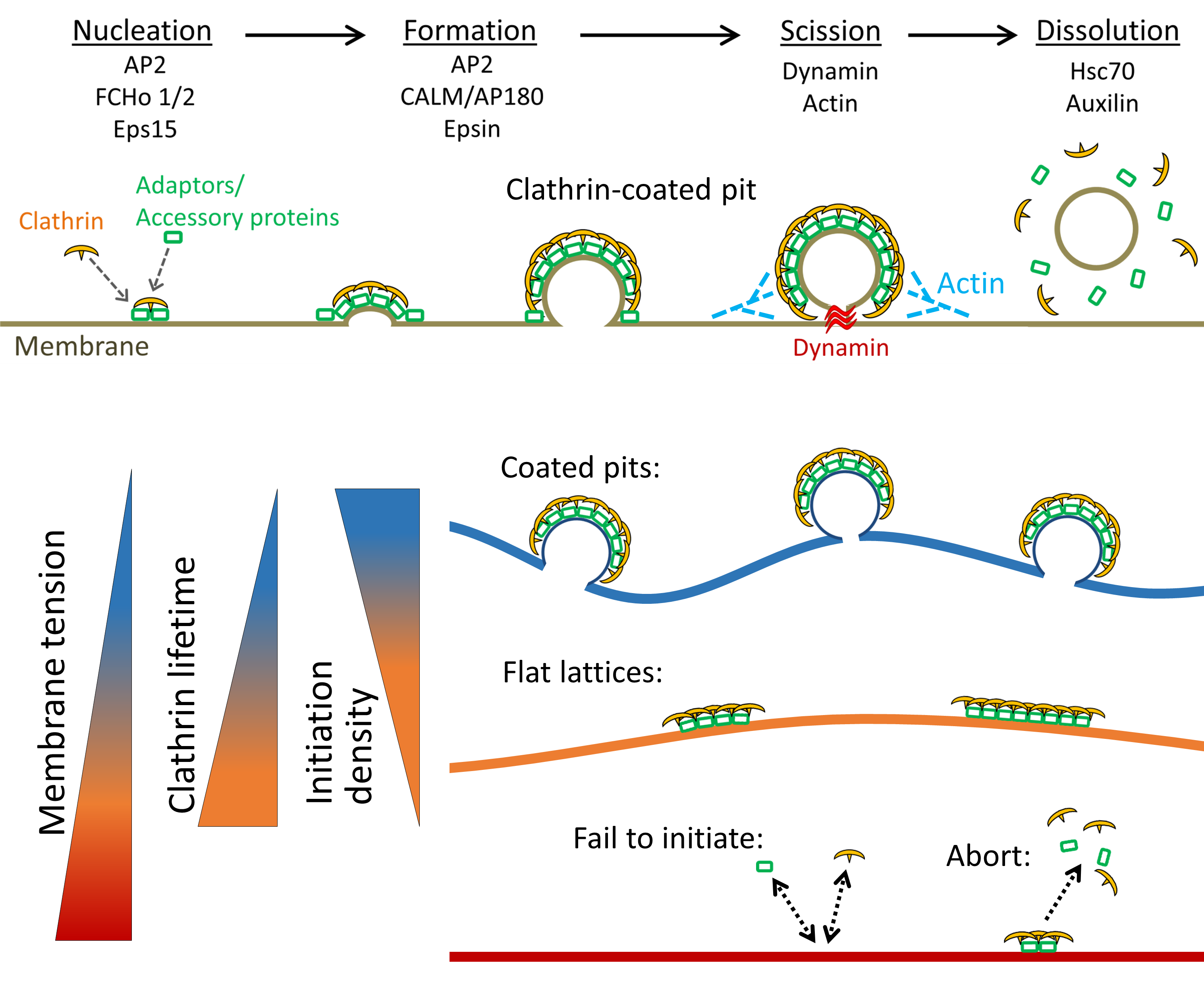
Dynamic interplay between cell membrane tension and clathrin-mediated endocytosis is published in Biology of the Cell.
2020

Emily is a Molecular Biophysics Training Program trainee now. Congratulations Emily.
2019

Caught by a pleasant surprise.

Dr. Umida Djakbarova has been awarded a Pelotonia Postdoctoral Fellowship. Congratulations Umida.
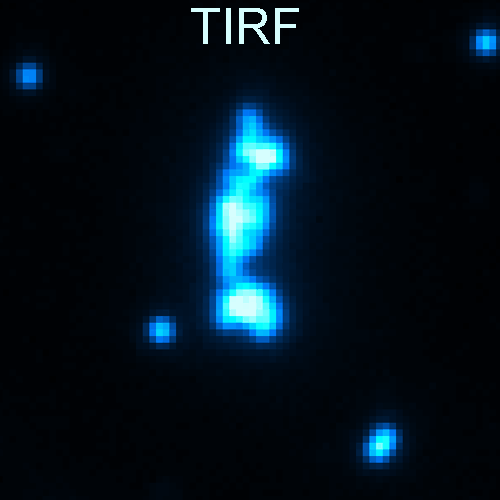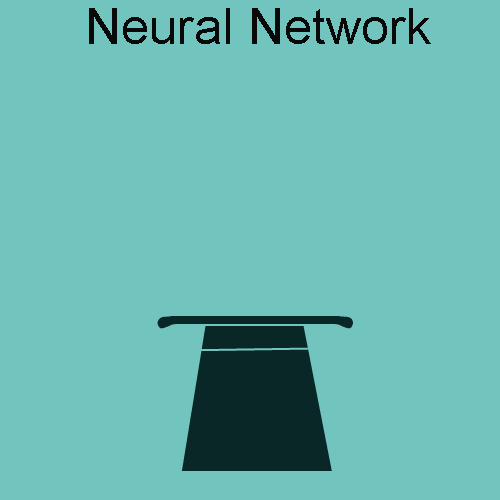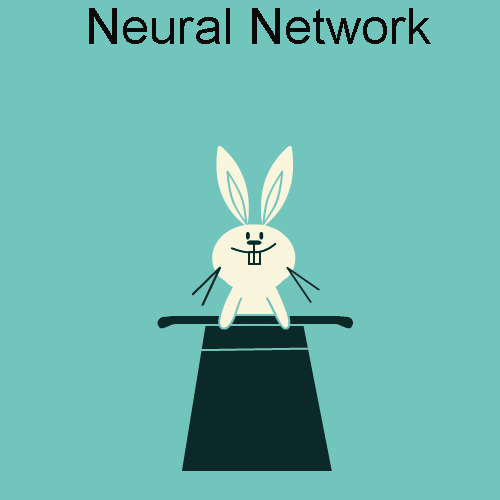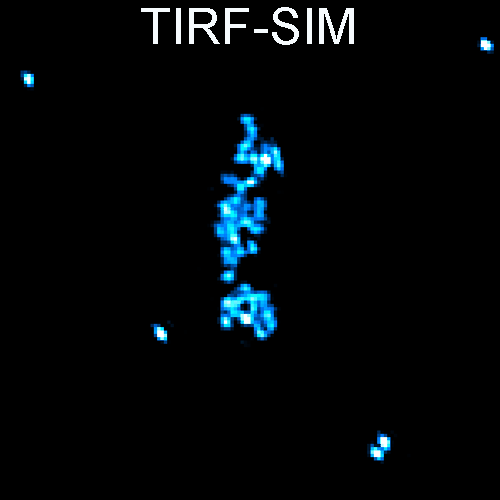
Deep learning enables cross-modality super-resolution in fluorescence microscopy is published in Nature Methods.

Scott has defended his dissertation. Congratulations Dr. Huber.
2018

Nathan has defended his dissertation. Congratulations Dr. Willy.

Josh has defended his dissertation. Congratulations Dr. Ferguson.
2017

Comert has received a National Science Foundation Early CAREER Development Award.

Joshua Ferguson has been awarded a Presidential Fellowship. Congratulations Josh.
We are on the cover of Molecular Biology of the Cell.

Nathan gave a platform presentation at the Fourth Midwest Membrane Trafficking and Signaling Symposium.

Our proposal to the Advanced Imaging Center has been accepted. We are going to the Janelia Research Campus.
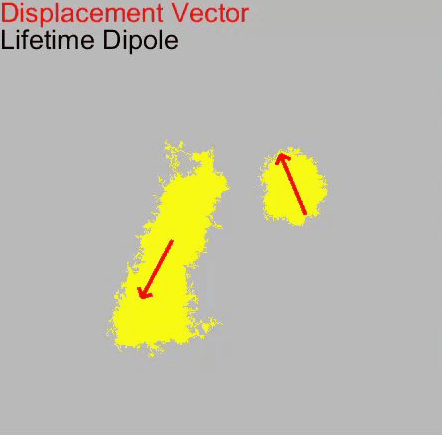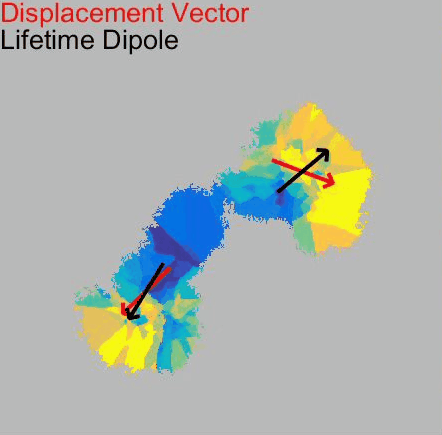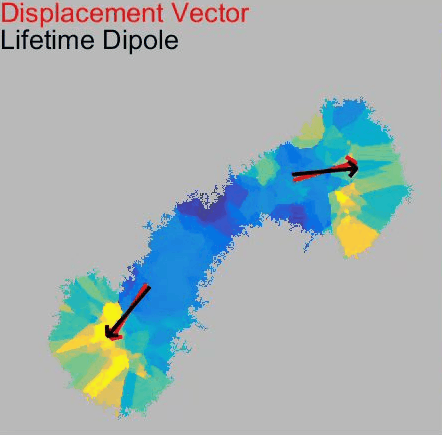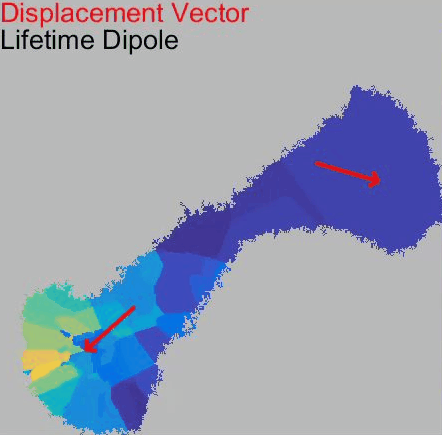
Our paper Membrane Mechanics Govern Spatiotemporal Heterogeneity of Endocytic Clathrin Coat Dynamics is now accepted by Molecular Biology of the Cell.
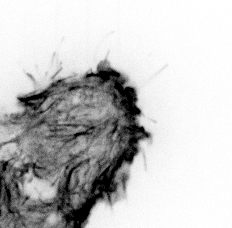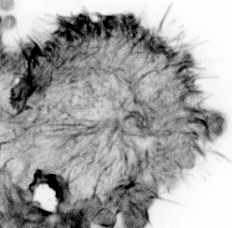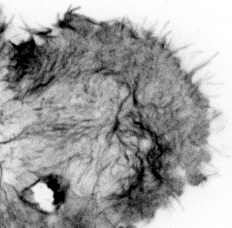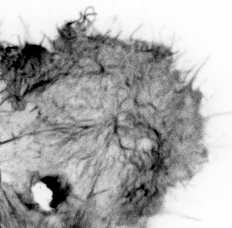
Our paper Mechanoregulation of Clathrin-mediated Endocytosis is now accepted by the Journal of Cell Science.
Josh gave a platform presentation at the Biophysical Society Meeting.
2016
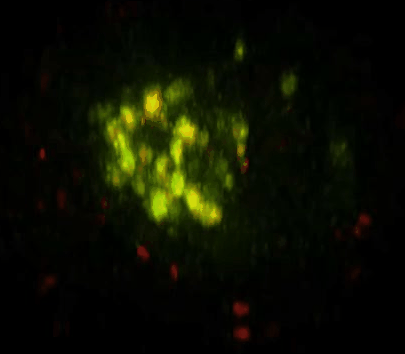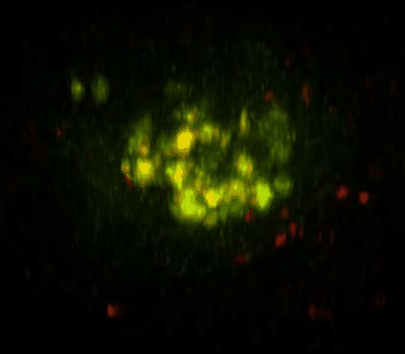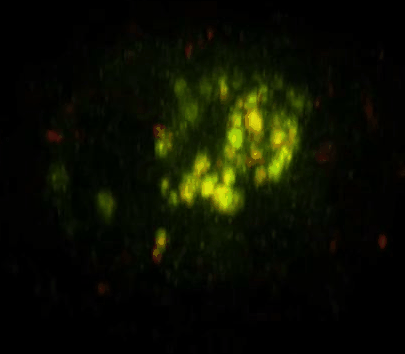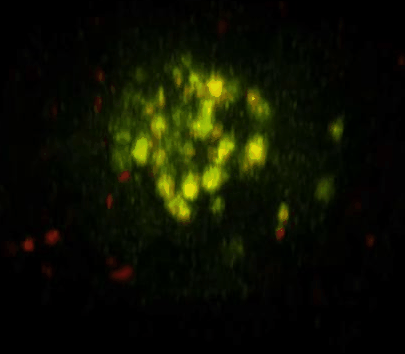
Daunorubicin-Loaded DNA Origami Nanostructures Circumvent Drug-Resistance Mechanisms in a Leukemia Model is published in Small.

Our paper Deciphering dynamics of clathrin-mediated endocytosis in a living organism is published in Journal of Cell Biology.
2015

Our paper Asymmetric formation of coated pits on dorsal and ventral surfaces at the leading edges of motile cells and on protrusions of immobile cells is published in Molecular Biology of the Cell.
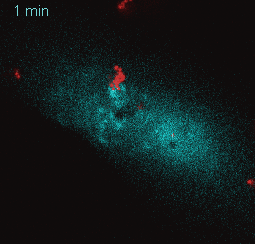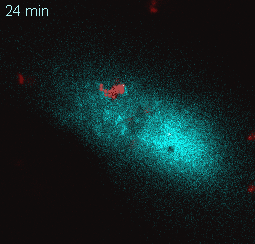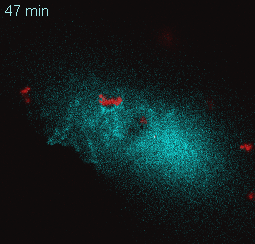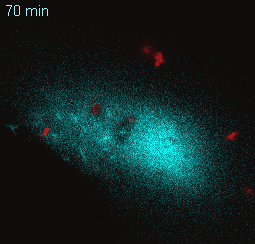
EtpE Binding to DNase X Induces Ehrlichial Entry via CD147 and hnRNP-K Recruitment, Followed by Mobilization of N-WASP and Actin is published in mBio.
2014

Spencer is accepted for the OSU 2014 Undergraduate Summer Research Scholarship.
2013
Matthew is selected to be a judge at The Ohio Academy of Science's State Science Day, encouraging students to pursue learning in science, engineering, technology, and education.
Our paper Similar uptake but different trafficking and escape routes of reovirus virions and infectious subvirion particles imaged in polarized Madin Darby canine kidney cells is published in Molecular Biology of the Cell.
Our spinning disk head has arrived.
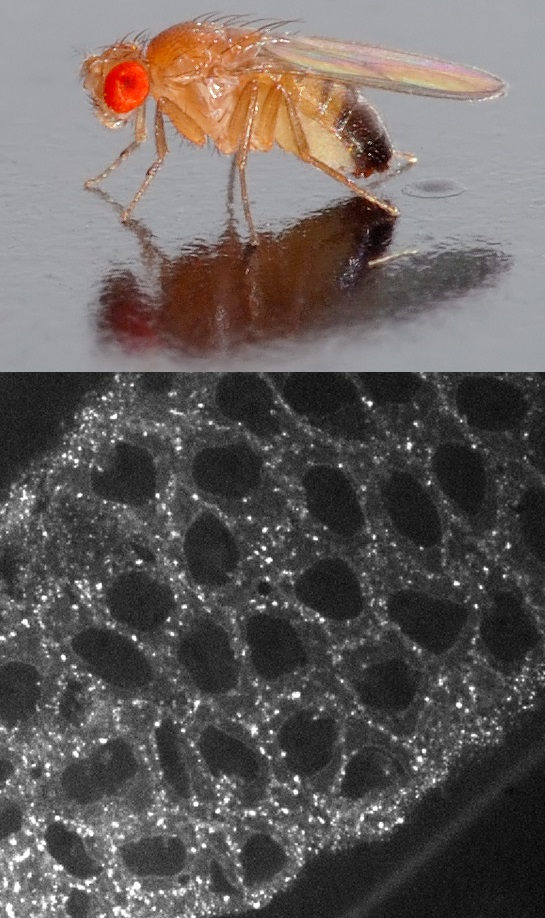
We are now officially culturing Drosophila melanogaster fly stocks and using our spinning-disk and light-sheet microscopy systems to image endocytic dynamics during embryogenesis.
2012
Matthew is awarded a Career Development Grant from the Ohio State University Council of Graduate Students to attend the 2013 annual meeting of the Biophysical Society in Philadelphia, Pennsylvania.
Our paper Dynamics of Intracellular Clathrin/AP1- and Clathrin/AP3-Containing Carriers is published in Cell Reports.
07 - Openings & Contact
We are always looking for highly motivated postdoctoral scholars, graduate students, and undergraduates with interests in cell biology, physics, or imaging. To apply, email Comert with your CV and a short note about your interests.